About the core and service
Mission
The Pharmacokinetics and Mass Spectrometry (PKMS) Core aims to facilitate researchers' efforts to discover new medicines, obtain research funding, file patent applications, and publish academic research findings.
About the Core
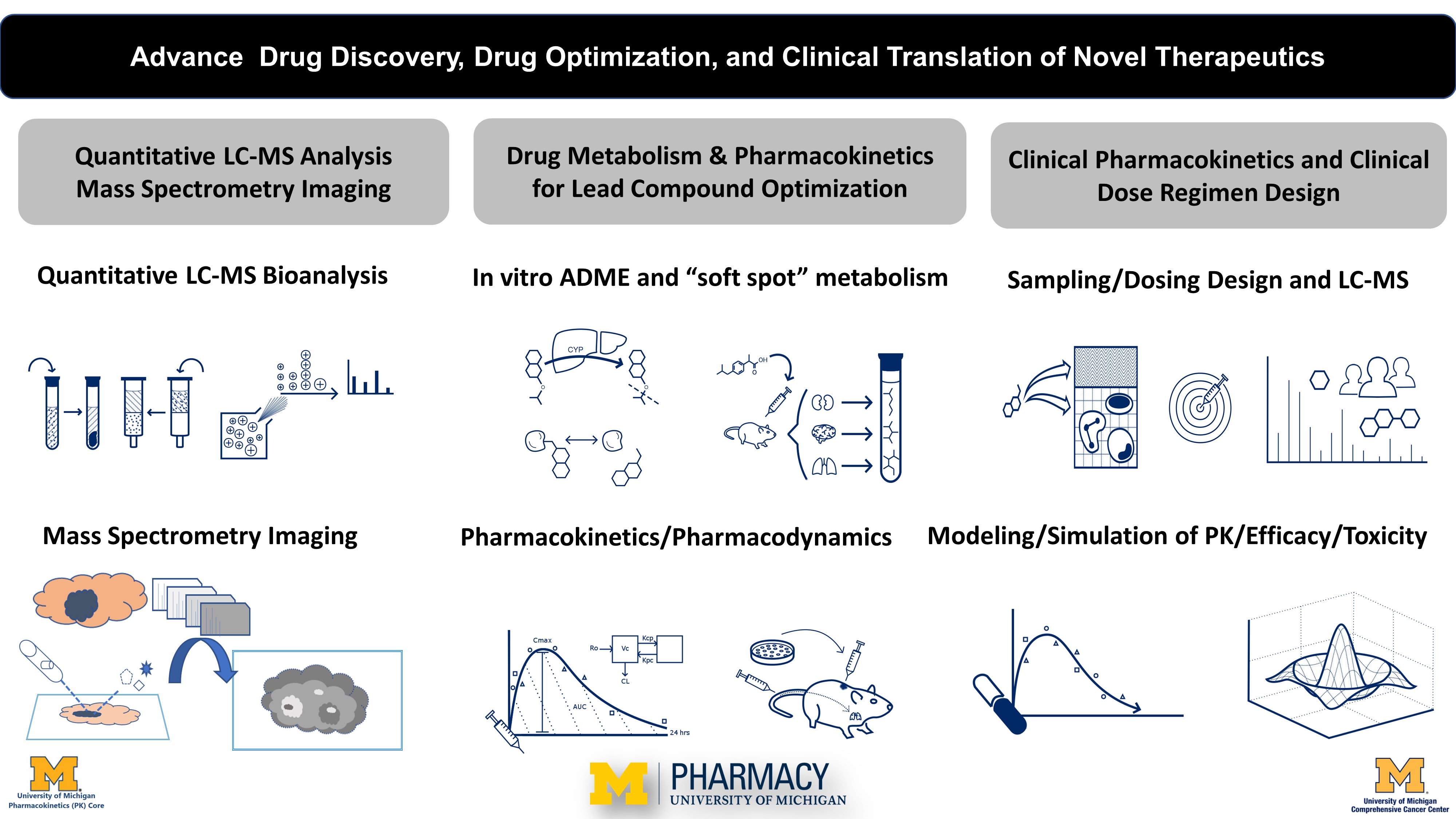
The pharmacokinetics and mass spectrometry (PKMS) core plays a pivotal role in advancing the drug discovery, clinical translation, and optimization of novel and existing therapeutics. The PKMS core has a thirteen year track record of supporting: (1) quantitative LC-MS analysis of molecules and mass spectrometry imaging of spatial localization biomarkers in tissue section; (2) preclinical ADME and pharmacokinetics for lead compound optimization in drug discovery and development; (3) clinical pharmacokinetics and dosage regimen design for clinical trials.
The PKMS core was founded in 2009 and has been funded by the cancer center supported grant (CCSG) since 2012. The PKMS core is led by Co-Directors Drs. Duxin Sun and Amit Pai. Dr. Sun has expertise in preclinical ADME, pharmacokinetics, drug discovery, and Nanomedicine. Dr. Pai has expertise in clinical pharmacokinetics and clinical trial design. Assistant director Dr. Bo Wen oversees the day-to-day operations that are run by ten full time staff members (four PhD and two MS level analytical chemists, two animal specialists, and two clinical study coordinators). PKMS core has eight LC-MS instruments that include three MS with high resolution MS capability and two MS with mass spectrometry imaging capability.
In the core's short history, we have supported LC-MS analysis, pre-clinical ADME and PK of more than 7500 compounds; supported clinical pharmacokinetics of more than 55 compounds in clinical trials. We have contributed to more than 310 grant applications, in which 101 grants were funded with a total funding of $260 million to the University of Michigan. We have provided services to more than 70 U-M labs, and 20 external labs. Our work has resulted in over 130 joint manuscripts, with five patents issued and six pending.
We can work with you either by joining your grant proposals as collaborators, for which we provide consultation as well as PK services, or fee-for-service via recharge. Please contact the PK Core assistant director, Dr. Bo Wen, if you are interested in working with us.
The PK Core is supported by the College of Pharmacy, the Comprehensive Cancer Center, and the Center for Discovery of New Medicines.
Flyer
Download our informational flyer:
Service
LCMS bioanalysis:
A. Advanced LC-MS/MS bioanalysis method development.
B. Method transfer and method qualification.
C. Quantitative analysis of small molecular and proteins from biological samples, or food and environment samples.
D. Drug and metabolite analysis to support PK/PD study on medicinal research.
E. Bioanalysis for clinical study by validated method according to FDA Guidance.
F. Chemical compound identification and structure elucidation.
In vitro/In vivo ADME and Preclinical Pharmacokinetics
A. Microsomal stability, Plasma stability, S9 stability, Hepatocyte stability,
B. Cytochrome P450 inhibition, Cytochrome P450 induction, Cytochrome P450 reaction phenotyping.
C. Plasma protein binding, Brain tissue binding, Blood to plasma ratio.
D. Metabolite identification and soft spot identification of target compounds.
E. Tissue bioanalysis including plasma and other organs from in vivo biodistribution study.
F. Oral bioavailability of drugs in rodent modes, or dogs.
G. Max tolerated dose with dose escalation protocol.
H. In vivo screening study with rodent to gain rapid insight into the compounds ADME behavior.
I. In vivo PK study with rodent or dogs to gain broader insight into the compounds PK profile.
J. Support customer designed animal PK study, including tumor mice model.
Clinical Pharmacokinetic studies
A. Clinical study development.
B. Data analysis with non-compartment model or non-linear mixed effects modeling.
C. Modeling and simulation with Population-PK analysis.
D. Precision medicine.
Advanced mass spectrometry application
A. Mass spectrometry imaging on tissue samples to visualize or provide spatial information of drugs and their metabolites, peptides, lipids, metabolisms, and proteins and their modification.
B. Biomarker identification and characterization.
C. Targeted metabolomics for bile acid, amino acid and glucose.
D. Targeted lipidomics.